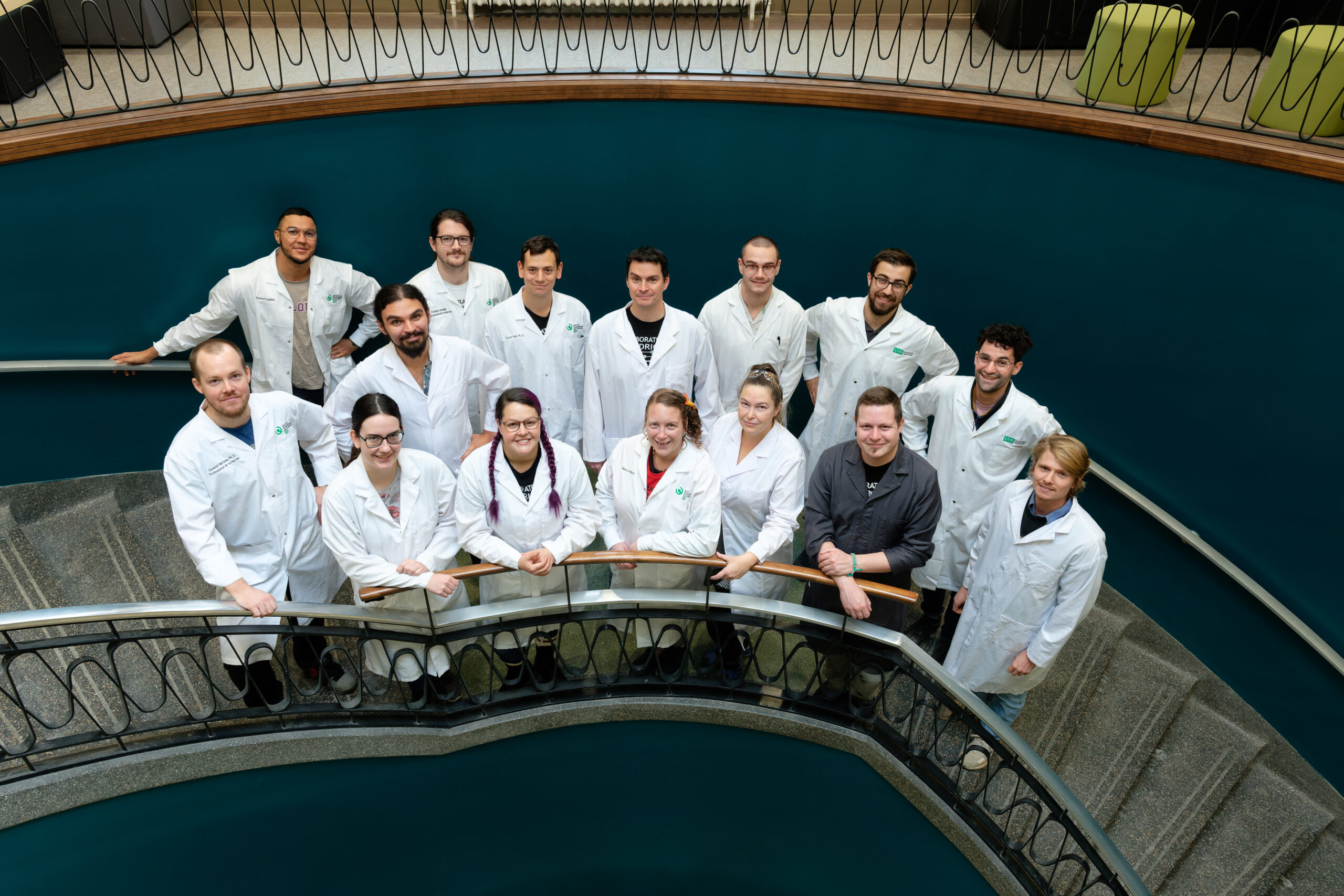
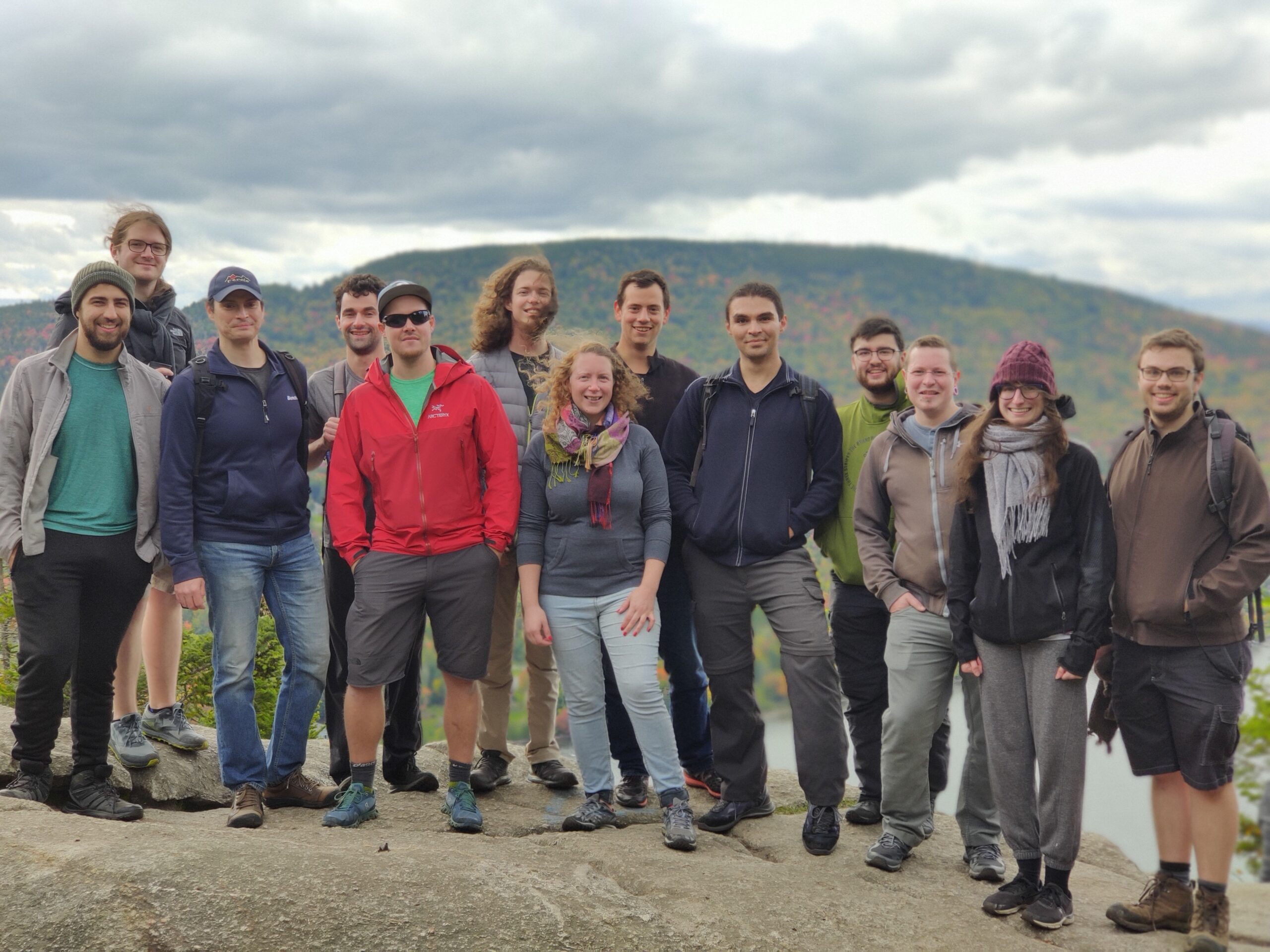
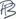Sebastien Rodrigue’s team, Université de Sherbrooke, December 2022
(credits: Michel Caron)
Principal Investigator
-
Sébastien Rodrigue, Ph.D.
Full Professor
TATUM Bioscience Co-Founder and CSO.
M.Sc. and Ph.D. Biologie, Université de Sherbrooke
Post-doctorate, Department of Civil and Environmental Engineering, Massachusetts Institute of Technology (MIT)
Postdoctoral fellows
-
Antoine Champie, Ph.D.
Investigating synthetic lethality in Escherichia coli.
-
Nancy Allard, Ph.D.
Developement of a minimal and predictable metabolic cellular chassis by the rational design and reduction of the Escherichia coli genome.
-
Kevin Neil, Ph.D.
TATUM Bioscience Co-Founder and CTO
Development of LBT-1 as an alternative technique to antibiotic
molecules.
-
Amel Ben Lagha, Ph.D.
(co-direction with Prof. Nathalie Rivard, Université de Sherbrooke)
Understanding the interplay between Shp-2 and microbiota in colonic
inflammation.
-
Alexandre Desroches, Ph.D.
(co-direction with Immune Biosolutions)
Graduate students
-
Simon Jeanneau, Ph.D. Student
(co-direction with Prof. Pierre-Étienne Jacques)
Integrative analysis of omics datasets and development of bioinformatic pipelines.
-
Nicolas Allaire-Tanguay, Ph.D. Student
Genetic engineering of conjugative plasmid TP114.
-
Julien Faure-Lévesque, Ph.D. Student
Development of bioinformatic tools for whole-genome design and assembly.
-
Jean-Christophe Berger Dancause, Ph.D. Student
(co-direction with Pr Daniel Lafontaine, Université de Sherbrooke)
-
David Pivin, Ph.D. Student
(co-direction with Pr François Michaud, Université de Sherbrooke)
Development of a robotic platform for synthetic biology.
-
Lachlan Bartrop, Ph.D. Student
(co-direction with Pr Louis-Patrick Haraoui, Université de Sherbrooke)
-
Anthony Duval, M.Sc. Student
Modification of the Mesoplasma florum genome.
-
Ryphed Lagdani, M.Sc. Student
Whole-genome synthesis, refactoring and assembly.
-
Amélie De Grandmaison, M.Sc. Student
Investigating synthetic lethality in Escherichia coli.
-
Jonathan Boisvert, M.Sc. Student
Development of an inducible expression system and a mimetic antibody secretion platform in the near-minimal bacterium Mesoplasma florum.
-
Jérémy Gagnon, M.Sc. Student
Construction of Mesoplasma florum synthetic genomes by cloning in Saccharomyces cerevisiae
-
Arrianne Pascale Collette, M.Sc. Student
(co-direction with Pr Jean-Philippe Côté, Université de Sherbrooke)
Undergraduate students
- Audrey Poirier
- Amélie Doucet
- Günes-Hélène Isitan (Diplôme d’études supérieures spécialisées en Sciences)
Research assistants
-
Frédéric Grenier, M.Sc.
-
Dominick Matteau, Ph.D.
Alumni
- Vincent Quévillon, undergraduate intern
- Willem Laverlochere, undergraduate intern
- Vincent Baby, post-doctoral student
- Patricia Roy, M.Sc. student
- Catherine Chamberland, M.Sc. student
- Olivier Trahan, M.Sc. student
- Samuel Côté, undergraduate intern
- Antoine Pelletier, undergraduate intern
- Jean-Christophe Lachance, Ph.D. student
- Romain Durand, Ph.D. student
- Jean-François Rousseau, M.Sc. student
- Cynthia Lemieux, M.Sc. student
- Marie-Eve Pepin, M.Sc. student
- David Jordan, undergraduate intern
- Ariane Arsenault, Cégep de Sherbrooke intern
- Alejandra Matsuri Rojano Nisimura, undergraduate intern (Mitacs Globalink)
- Yogesh Lakhotia, undergraduate intern (Mitacs Globalink)
- Jimmy Blouin, undergraduate intern
- Samuel Jacques, undergraduate intern
- Lucas Leclerc, undergraduate intern
- Charles Coulombe, undergraduate intern
- Mélissa Arango-Giraldo, undergraduate intern
- Samuel Gauthier, undergraduate intern
- Syed Fazle Rouf, post-doctoral student
- Joëlle Brodeur, research assistant
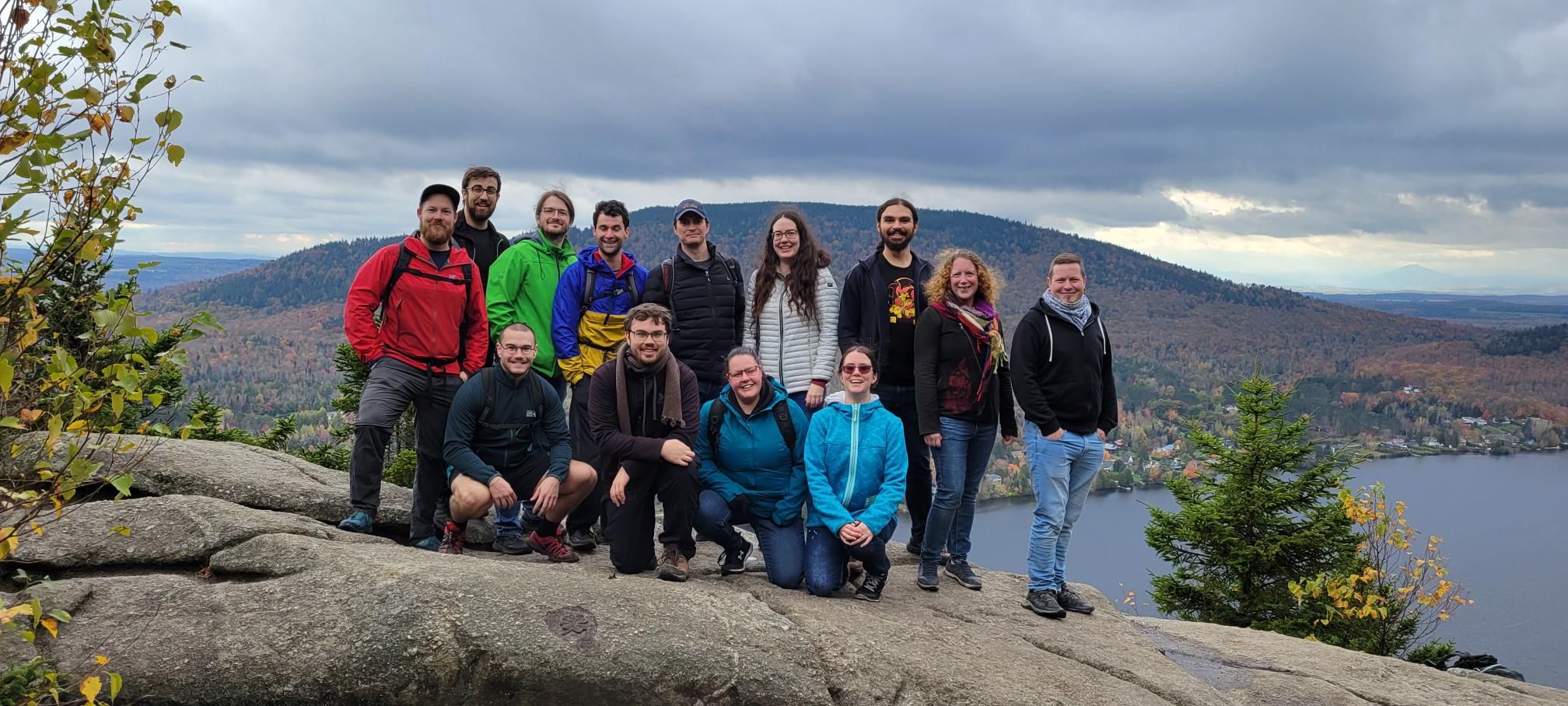
Mount Pinacle, Coaticook (Québec), October 2023
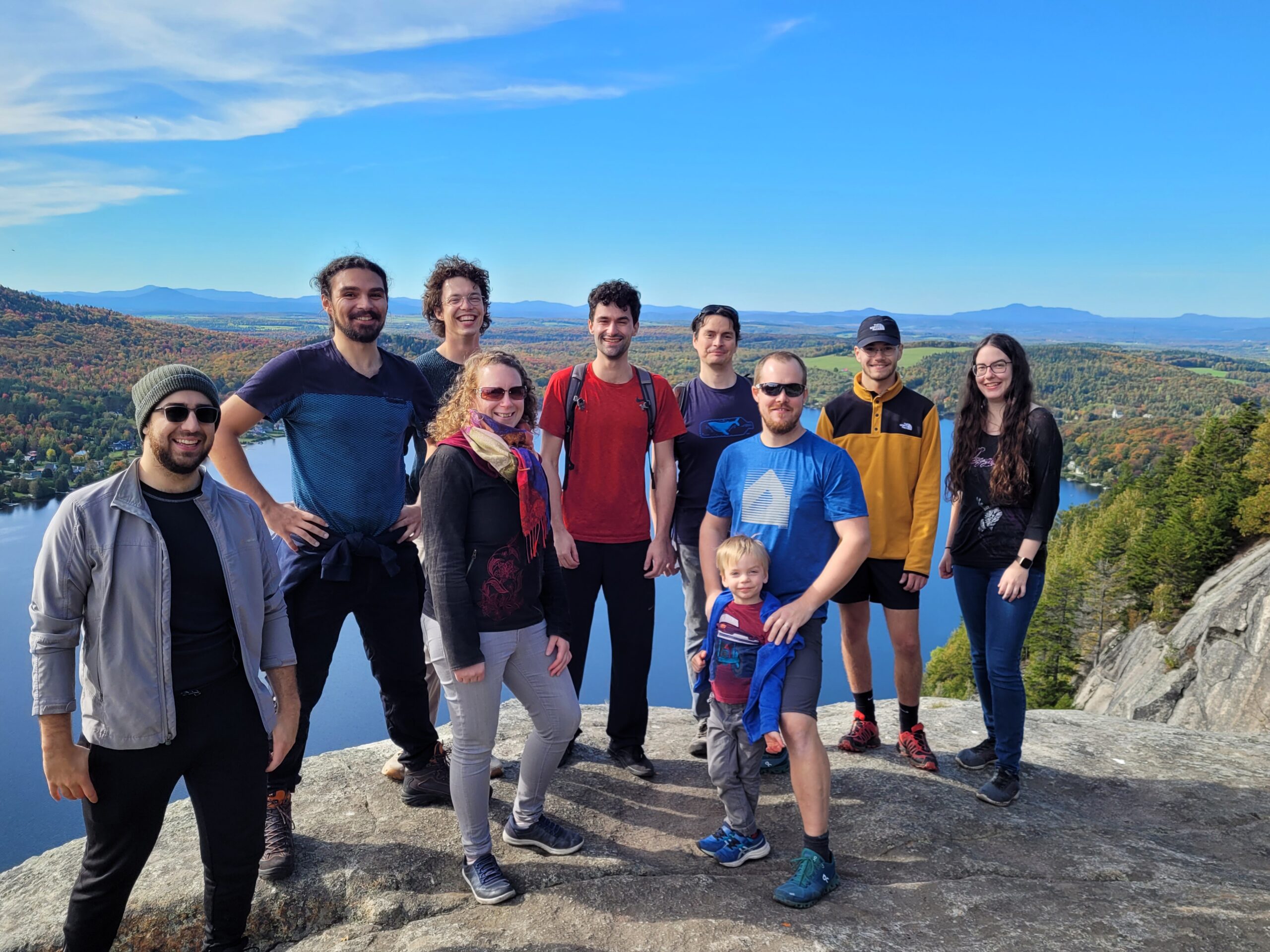
Mount Pinacle, Coaticook (Québec), October 2022
Mount Pinacle, Coaticook (Québec), October 2021
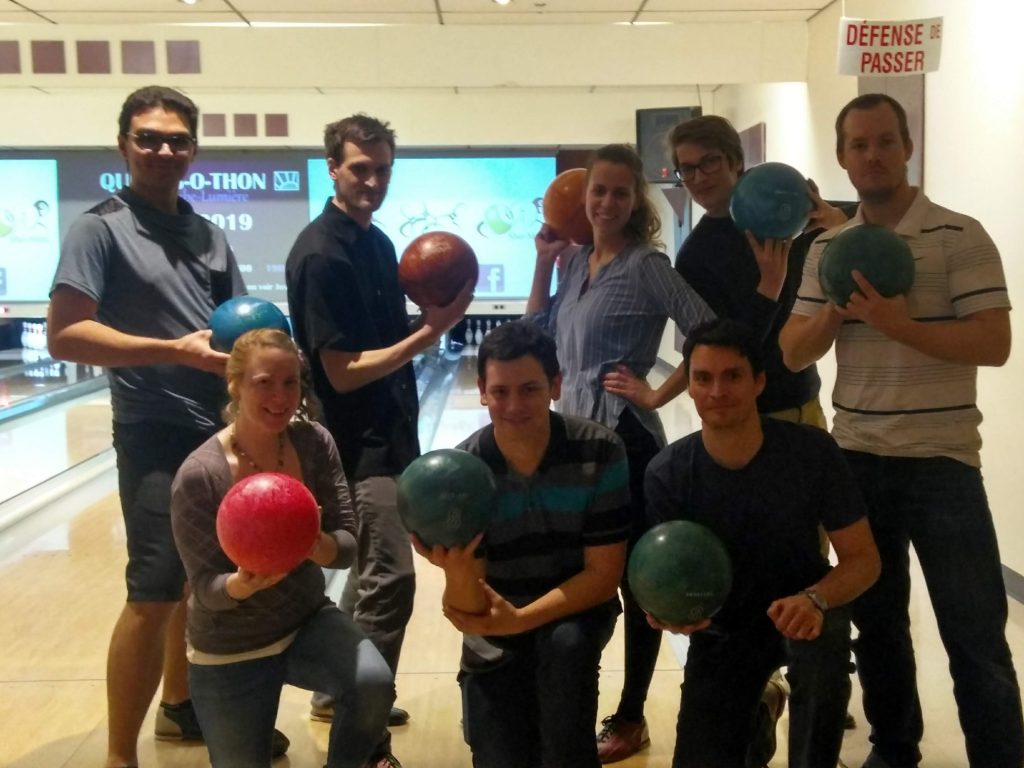
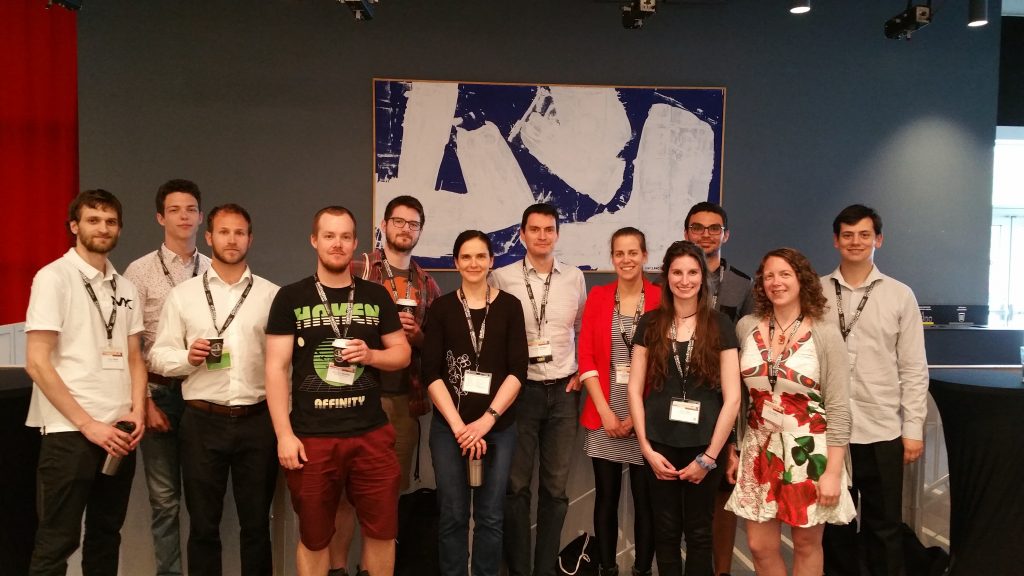


69th Annual Conference of the Canadian Society of Microbiologists, June 2019